Country |
Genotype status |
n |
Allele frequency |
References |
||
|
D/D |
D/O |
|
O/O |
|
|
Burundi |
7 |
2 |
2 |
11 |
16/22=0.73 |
12 |
Cameroon 1 |
15 |
17 |
4 |
36 |
47/72= 0.65 |
8 |
Cameroon 2 |
90 |
53 |
30 |
173 |
233/346= 0.67 |
10 |
Cameroon 3 |
||||||
a) Bamiléké |
67 |
102 |
44 |
213 |
236/426= 0.55 |
13 |
b) Other ethnies |
140 |
39 |
16 |
195 |
319/390= 0.82 |
13 |
c) All ethnies |
207 |
141 |
60 |
408 |
555/816= 0.68 |
13 |
CAR |
0 |
5 |
3 |
8 |
5/16= 0.31 |
7, this work |
DRC |
20 |
18 |
0 |
38 |
58/76= 0.76 |
This work |
RC |
0 |
1 |
1 |
2 |
1/4= 0.25 |
This work |
South Africa |
53 |
25 |
7 |
85 |
131/170= 0.77 |
5,7 |
Tanzania |
7 |
6 |
0 |
13 |
20/26= 0.77 |
7 |
Zambia |
5 |
1 |
1 |
7 |
11/14= 0.79 |
7 |
Zimbabwe |
27 |
4 |
0 |
31 |
58/62= 0.94 |
8,9 |
JSM Dermatology and Clinical Research
-
Research ArticleWhat can we learn from the Distribution of the 2.7kb Deletion Mutation of the OCA2 Gene in Oculocutaneous Albinism Type 2 (OCA2) in Cameroon and in Sub-Saharan Countries?Robert Aquaron1*, Jean-Louis Berge-Lefranc2, Eulalie Lassaux3, Claudio Plaisant3, Benoit Arveiler3,5 and Murray Brilliant41Timone Health Campus, Aix-Marseille University, France
2Laboratory of Molecular Biology, Conception Hospital, France
3Department of Medical Genetics, Bordeaux University Hospital, France
4Center for Precision Medicine Research, Marshfield Clinic Research Institute, USA
5University of Bordeaux, Rare Diseases: Genetics and Metabolism (MRGM), France*Corresponding author: Robert Aquaron, Timone Health Campus, Aix- Marseille University, Boulevard Jean Moulin, 13385, Marseille, France, Tel: 33-06-1374-0767; Email: robert.aquaron@univ-amu.frSubmitted: 08 February 2019; Accepted: 18 April 2019; Published: 19 April 2019 -
The specific African intragenic deletion of the 2.7-kb of the OCA2 gene was determined in a 408 OCA2 patients from Cameroon with an allele frequency of 0.68, a value close to that found in other Bantu-speaking countries located in central, south and east Africa. We observed a lack of this mutation in 30 non-Bantu-speaking OCA2 patients from West Africa countries. This mutation is associated in Cameroonian OCA2 subjects with a specific TAG haplotype allowing us to estimate that this mutation originated 4,100-5,645 years ago probably in the original Bantu homeland situated in the present day at the border Nigeria-Cameroon. The eastern wave of expansion and migration of Bantu-speaking peoples from their homeland was studied by comparing haplotype analysis using 4 intragenic microsatellites markers of OCA2 subjects from Cameroon and Democratic Republic of Congo. The haplotype 232-156 was present at frequency 1.0 in Cameroonian OCA2 patients and 0.87 in OCA2 DRC subjects suggesting that this haplotype is originating from Cameroon and was diluted during the migration process by genetic contribution of neighbouring populations. Finally we show that when this mutation was at homozygous state, the skin dendritic freckles were always absent, characterizing the phenotype OCA2a.Keywords: OCA2; 2.7kb deletion; Sub-Saharan Africa; Cameroon; Bantu
-
OCA: Oculo Cutaneous Albinism; TYR: Tyrosinase, TYRP1: Tyrosinase Related Protein 1; SLC: Solute Carrier; P: Pinkeye dilution; ER: Endothelial Reticulum; CAR: Central African Republic; DRC: Democratic Republic of Congo; RC: Republic of Congo; SNP: Single Nucleotide Polymorphism; DF: Dendritic FrecklesOculocutaneous Albinism (OCA) is a rare genetic disease characterized by generalized hypopigmentation of the skin, hair and eyes causing multiple ophthalmologic abnormalities (decreased visual acuity, nystagmus, light sensitivity) and impaired eye development (incomplete development of the fovea and changes in axonal routing in the optic nerve and tracts). OCA is a genetically heterogeneous group of autosomal recessive disorders caused by deficiency of melanin biosynthesis or melanin transport. Melanin is a polymeric pigment produced in specialized ectodermally derived cells called melanocytes and in organelles called melanosomes. There are four common forms of non-syndromic OCA in which the manifestations are limited to skin, hair and eye function: OCA1, OCA2, OCA3 and OCA4 named after their date of the mutations discovered respectively in genes TYR, OCA2, TYRP1 and SLC45A2. OCA2 is caused by mutations in the OCA2 gene (formerly called P gene) on Chromosome 15q12 that contains 25 exons. The encoded protein is predicted to have a 12-transmembrane domains and functions as a chloride transporter [1]. However disruption of the function of P gene causes the aberrant trafficking of TYR, the key enzyme required for melanin synthesis, to melanosomes after its processing through the ER and Golgi [2].OCA2 is the most common type of OCA among Africans and African-Americans. There is a high incidence of OCA2 among specific African populations mainly in sub-Saharan countries located in south, central and east Africa. A specific intragenic deletion of 2.7kb of the OCA2 gene lacking exon 7 has been described in 1994 [3]. This mutation, at homozygous or heterozygous state, was found in four unrelated African-American OCA2 patients, in two African OCA2 patients from Cameroon and DRC, but not in Caucasians with OCA2. This finding indicates an African origin for this allele and was confirmed in several studies [4-13].The aims of this work are a) to recapitulate our research on the frequency of this mutation mainly in Cameroon and in some other African countries (Burundi, DRC, RC) in comparison with the findings in other African countries, b) to postulate that the haplotypes associated with this deletion could be utilized as a biological marker for the migration of Bantu-speaking people and c) that individuals who are homozygous for this mutation correspond to the subtype, OCA2a, characterized by the absence of dendritic freckles.We analyzed 459 OCA2 patients from different countries (Cameroon, CAR, DRC, RC, Burundi) for the 2.7kb deletion study. We analyzed 27 OCA2 patients from Cameroon and 38 from DRC for the haplotype study. We analyzed 124 OCA2 patients from Cameroon and Tanzania for the OCA2a and OCA2b study. Informed written and/or oral consent was obtained from the patients. For each patient a clinical data sheet with skin and ophthalmologic findings was performed. PCR analysis of the 2.7 kb deletion was performed using either semi-quantitative PCR or qualitative PCR [14] Haplotype analysis of the OCA2 locus was performed using four intragenic microsatellite markers located respectively in Introns 2, 17, 20 and 22: OCAI-2, OCAI-17, OCAI- 20 and OCAI-22a) Since 1973, we have been focused on understanding OCA in Cameroon and particularly by OCA2 which is the most frequent type of albinism in this country, in Sub-Saharan countries and in African-Americans resulting from the Atlantic slave trade. Since the discovery in 1994 [3] of the specific African intragenic deletion of 2.7-kb of the OCA2 gene in OCA2 Africans (Cameroon, DRC) and in African-Americans, we have been able to easily detect this specific mutation at homozygous or heterozygous state or its absence among OCA2 patients from Cameroon and other Sub-Saharan countries (Burundi, CAR, DRC and RC). The genotypic status and the allele frequency of this mutation are summarized in Table 1. Among a total of 730 OCA2 Cameroonian patients examined, we performed analysis of the deletion allele respectively in 36, 173 and 408 individuals showing respectively an allele frequency of 0.65 [8]; 0.67 [10] and 0.68 [13]. In the 408 patients study we reported separately results from the principal ethnic group, the Bamileke tribe with an allele frequency of 0.55 among 213 subjects [13]. Off the 195 individuals of different ethnic groups (Bamoun, Bassa, Beti, Douala, Ewondo) the allele frequency was 0.80 resulting in a median value of 0.68 for Cameroon. As a whole the allele frequency in some central, south and east Africa countries was around 0.70 (Cameroon: 0.68; Burundi: 0.73; DRC: 0.76; South Africa and Tanzania: 0.77; Zambia: 0.79) culminating with 0.94 in Zimbabwe (Figure 1). The low values found in CAR (0.31) and in RC (0.25) may result from the paucity of analyzed patients 2 and 8 respectively. This mutation was also found in African- Americans from USA and Haiti [3,4] with a low allele frequency 7/24= 0.29 due to the low number of subjects (12) but also due to the fact that the Atlantic slave trade has occurred mainly ( >55%) from West Africa where the population is predominantly negroid [15]. These findings were in accordance with our results: a lack of this mutation in 30 non Bantu-speaking OCA2 individuals from West African countries: 17,10,1,1 and 1 respectively from Mali , Niger, Nigeria, Togo and Burkina-Faso. Another child with OCA2 from Nigeria lacking this mutation has also been reported [6]. The high frequency of the deletion allele could be due to a selective advantage enjoyed by the heterozygotes or as a result of genetic drift or by some unknown selective force. It has also been hypothesised that carriers of this mutation may have a protective advantage from infection with leprosy [16]. This mutation was associated predominantly in Southern Africa [7] with one common haplotype determined by Southern blot of genomic DNA (A: 0.78). Subsequently using a five SNP assay we identified in 53 OCA2 Cameroonian [11] a common TAG haplotype with frequency of 1.0. This study allowed us to estimate that this mutation originated 4,100- 5,645 years ago. These findings showed that this mutation was specific of Bantu speaking populations living in central, south and east Africa. Anthropological and archaeological data seem to indicate that original Bantu homeland is situated in the present day at border Nigeria-Cameroon near the Benoue River [17]. Linguistic and genetic data fit well with a scenario of dispersal and expansion of Bantu from their homeland around 5,000 years ago. Genetic data were based on uniparental markers: mtDNA for females and Y chromosomes for males [18-20]. The expansion of Bantu speaking peoples occurred in two directions: the southernwestern wave moved mostly through the coast, the riverine sites then to east; the eastern wave involved the settlement of farmers in a region which is now Uganda, then to the great lake region, then to south and west (Figure 2). The two waves of expansion are thought to have met around 2.000 years ago, in the region now known as Zimbabwe. Today the migration process is still in progress. We had the opportunity to analyze a Bantu family with 2 children with OCA2 carrying the 2.7-kb mutation at homozygous state in Reunion island [21]. Reunion Island is a French overseas “department” located in the western Indian Ocean, off the eastern coast of Madagascar. In fact this family was originated from Mayotte, an island from the Comoros archipelago located northwest of Madagascar and probably from east Africa coast [22] (Figure 2).
-
 Table 1:Genotype status and allele frequencies of the 2.7-kb deletion of the OCA2 gene in OCA2 patients from different countries in east, south and
central Africa (n: number of individuals), D, Deletion mutation, O, Other mutation, CAR, Central African Republic, DRC, Democratic Republic of Congo,
RC, Republic of Congo. View Table
Table 1:Genotype status and allele frequencies of the 2.7-kb deletion of the OCA2 gene in OCA2 patients from different countries in east, south and
central Africa (n: number of individuals), D, Deletion mutation, O, Other mutation, CAR, Central African Republic, DRC, Democratic Republic of Congo,
RC, Republic of Congo. View Table
-
 Figure 1: Frequency of the 2.7-kb deletion allele of the OCA2 gene in
OCA2 patients from different sub-Saharan countries in east, central,
south and west Africa. View Figure
Figure 1: Frequency of the 2.7-kb deletion allele of the OCA2 gene in
OCA2 patients from different sub-Saharan countries in east, central,
south and west Africa. View Figure
-
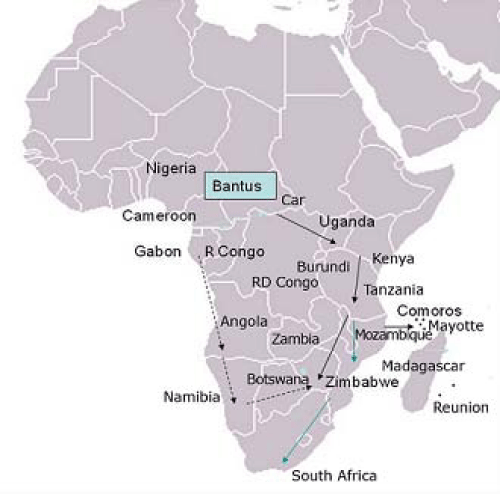 Figure 2:Bantu homeland, migration and expansion. Bantu tag refers
to the assumed original Bantu area at border Nigeria- Cameroon near
the Benoue River. Filled arrows refer to the eastern Bantu waves
with settlement of farmers in the interlacustrine region which is now
Uganda. Then the expansion was directed south in three distinct
waves, one travelling along the coast (blue arrows) , the second going
to Comoros archipelago and Madagascar island and the third one
going through west. Dotted arrows refer to the southward western
waves moving through the coast then east. The two major waves
(filled and dotted arrows) are thought to have met in the region now
known as Zimbabwe (adapted from [18,19]). View Figure
Figure 2:Bantu homeland, migration and expansion. Bantu tag refers
to the assumed original Bantu area at border Nigeria- Cameroon near
the Benoue River. Filled arrows refer to the eastern Bantu waves
with settlement of farmers in the interlacustrine region which is now
Uganda. Then the expansion was directed south in three distinct
waves, one travelling along the coast (blue arrows) , the second going
to Comoros archipelago and Madagascar island and the third one
going through west. Dotted arrows refer to the southward western
waves moving through the coast then east. The two major waves
(filled and dotted arrows) are thought to have met in the region now
known as Zimbabwe (adapted from [18,19]). View Figure
b) In addition to the classical genetic markers (mtDNA and Y chromosomes) utilized for the study of Bantu migration, we have shown that the frequency of the 2.7-kb deletion allele is around 0.70 in the homeland of Bantus, Cameroon and in countries located on the eastern wave: DRC, Burundi, Tanzania, Zambia and South Africa. The fact that Zimbabwe presented the highest allele frequency rate of 0.94 could be in accordance with the postulated meeting point of the 2 waves. We hypothesised that this mutation and new associated haplotypes in two of these countries (Cameroon and DRC) could be a marker of the eastern wave. We performed a haplotype analysis encompassing the OCA2 gene using four intragenic micro satellite markers: OCAI- 2, OCAI-17, OCAI-20 and OCAI-22. The haplotype 232-156-193- 147 was the only haplotype associated with the 2.7-kb deletion in Cameroonian albinos. The OCAI-2-OCAI-17 haplotype 232- 156 was present at frequency of 1.0 (44/44 alleles) in 22 OCA2 Cameroonian and 0.87 (58/68 alleles) in 34 OCA2 Congolese (DRC). In DRC albino beside this haplotype other OCAI-2-OCAI-17 haplotypes were found: 230-162, 230-168. These differences were probably due to different genetic contributions from neighbouring populations during the migration process.c) OCA2 individuals in southern Africa presented with two distinct phenotypes: affected individuals are characterized by the presence or absence of pigmented patches (ephelides) mainly on sun exposed areas [5]. The correlation between the 2 phenotypes and the deletion allele frequency showed a significant difference using the khi2 test (P<.0001): 0.89 (n: 24) for individuals without ephelides and 0.54 (n: 38) for those with ephelides. These pigmented skin lesions have been later clinically carefully described as dendritic freckles characterized by irregular branched shape, light to brown- black colour and large size (0.5 to 3.0cm). They appear only at the age 5-10 years [23]. These two phenotypes are now called: OCA2a without freckles and OCA2b with such freckles [24]. In a recent work we have found different results in a large cohort of 124 OCA2 individuals from Cameroon and Tanzania [25]. When the 2.7- kb deletion was present at homozygous state (84 OCA2 patients) DF were always absent. When the 2.7-kb deletion was at heterozygous state and the second mutation was a nonsense mutation like R165X, DF was also absent (3 Cameroonian patients). If the second mutation is a missense mutation like P221L we observe DF. We could hypothesise that when the mutations are deletions or nonsense, inactivation of the gene was total and irreversible and DF was always absent. In the case of missense mutations, the mutation can mutate back to normal in some melanocytes with sun damage leading to DF. This hypothesis is in good correlation with the fact that DF appears only at age 8-10 after sun exposure which is particularly important in Sub-Saharan Africa.We have shown that the specific African intragenic deletion of 2.7-kb of the OCA2 gene is present at a high allele frequency, around 0.70 in OCA2 individuals in many countries from east, south and central Africa but not in west Africa. It could be utilized as a genetic marker to follow the expansion and migration of Bantus from their homeland Cameroon, to south and east African countries. We have also shown that this mutation at homozygous state characterized the phenotype OCA2a where the skin dendritic freckles are absent.We thank all the volunteers from Cameroon, Burundi, DRC and Tanzania who participated in this study, Dr. Frederic Ntwengarubum from Bujumbura Burundi, Pr. Leon Tshilolo and Mwimba Texas from Kinshasa, DRC, Servain and Jeanjanvier Ndumba from Kisangani, DRC, Luc Kamdem from Yaounde and Hugues Ngando from Douala, Cameroon, for help with patients recruitment and blood samples supply. We also thank Ghislaine Hancy for help to preparing the manuscript. This work was supported by Genespoir (French Albinism Association). -
-
- Bellono NW, Escobar IE, Lefkovith AJ, Marks MS, Oancea E. An intracellular anion channel critical for pigmentation. ELife. 2014; 3: e4543.
- Toyofuku K, Valencia JC, Kushimoto T, Costin GE, Virador VM, Vieira WD, et al. The etiology of oculocutaneous albinism (OCA) type II: the pink protein modulates the processing and transport of tyrosinase. Pigment Cell Res. 2002; 15: 217-224.
- Durham-Pierre D, Gardner JM, Nakatsu Y, King RA, Francke U, Ching A, et al. African origin of an intragenic deletion of the human P gene in tyrosinase positive oculocutaneous albinism. Nat Genet. 1994; 7: 176-179.
- Lee ST, Nicholls RD, Schnur RE, Guida LC, Lu-Kuo J, Spinner NB, et al. Diverse mutations in the P gene among African-Americans with type II( tyrosinase- positive) oculocutaneous albinism (OCA2). Hum Mol Genet. 1994; 3: 2047-2051.
- Stevens G, van Beukering J, Jenkins T, Ramsay M. An intragenic deletion of the P gene is the common mutation causing tyrosinasepositive oculocutaneous albinism in southern African negroids. Am J Hum Genet. 1995; 56: 586-591.
- Spritz R, Fukai K, Holmes A, Luande J. Frequent intragenic deletion of the P gene in Tanzanian patients with type II oculocutaneous albinism (OCA2). Am J Hum Genet. 1995; 56: 1320-1323.
- Stevens G, Ramsay M, Jenkins T. Oculocutaneous albinism (OCA2) in sub-Saharan Africa: distribution of the common 2.7-kb P gene deletion mutation. Hum Genet. 1997; 99: 523-527.
- Puri N, Durham-Pierre D, Aquaron R, Lund PM, King RA, Brilliant MH. Type 2 oculocutaneous albinism (OCA2) in Zimbabwe and Cameroon: distribution of the 2.7-kb deletion allele of the P gene. Hum Genet. 1997; 100: 651-656.
- Lund PM, Puri N, Durham-Pierre D, King RA, Brilliant MH. Oculocutaneous albinism in an isolated Tonga community in Zimbabwe. J Med Genet. 1997; 34: 733-735.
- Aquaron R, Berge-Lefranc JL. Type 2 oculocutaneous albinism in Cameroon: distribution of the 2.7-kb deletion allele of the P gene among various ethnic groups. Pigment Cell Res. 2002; 9: 63.
- Aquaron R, Soufir N, Berge-Lefranc JL, Badens C, Austerlitz F, Grandchamp B. Oculocutaneous albinism type 2 (OCA2) with homozygous 2.7-kb deletion of the P gene and sickle cell disease in a Cameroonian family. Identification of a common TAG haplotype in the mutated P gene. J Hum Genet. 2007; 52: 771-780.
- Aquaron R, Berge-Lefranc JL, Grandchamp B. What can we learn from mutations in the oculocutaneous type II (OCA2) gene about African migrations “in” and “out” of Africa? Pigment Cell Melanoma Res. 2008; 21: 262.
- Aquaron R, Berge-Lefranc JL, Djatou M, Kamdem l. L’albinisme oculocutané dans le pays Bamileke, Cameroun. Sci Med Afr. 2011; 3: 362-374.
- Lasseaux E, Plaisant C, Michaud V, Pennamen P, Trimouille A, Gaston L, et al. Molecular characterization of a series of 990 index patients with albinism. Pigment Cell Melanoma Res. 2018; 31: 466-474.
- Salas A, Carracedo A, Richards M, Macaulay V. Charting the ancestry of African Americans. Am J Hum Genet. 2005; 77: 676-680.
- Tuli AM, Valenzuela RK, Kamugisha E, Brilliant MH. Albinism and disease causing pathogens in Tanzania: are alleles that are associated with OCA2 being maintained by balancing selection? Med Hypotheses. 2012; 79: 875-878.
- Berniell-Lee G, Calafell F, Bosch E, Heyer E, Sica L, Mougiama-Daouda P, et al. Genetic and demographic implications of the Bantu expansion: insights from human paternal lineages. Mol Biol Evol. 2009; 26: 1581-1589.
- Pereira L, Gusmao L, Alves C, Amorin A, Prata MJ. Bantu and european Y-lineages in Sub-Saharan Africa. Ann Hum Genet. 2002; 66: 369-378.
- Beleza S, Gusmao L, Amorin A, Carrado A, Salas A. The genetic legacy of western Bantu migrations. Hum Genet. 2005; 117: 366-375.
- Castri L, Tofanelli S, Garagnani P, Bibi C, Fosella X, Pelotti S, et al. mtDNA variability in two bantu-speaking populations (Shona and Hutu) from eastern Africa: implications for peopling and migrations patterns in sub-Saharan Africa. Am J Phys Anthropol. 2009; 140: 302-311.
- Aquaron R, Berge-Lefranc JL, Badens C, Roche J, Fite A, Sainte-Marie D, et al. L’albinisme oculocutané dans les départements français d’Outremer (Réunion, Guyane, Martinique) et à Mayotte : à propos de 21 observations dans 16 familles. Med Trop. 2005; 65: 584-591.
- Dubut V, Murail P, Pech N, Thionville MD, Cartault F. Inter- and extraindian admixture and genetic diversity in Reunion island revealed by analysis of mitochondrial DNA. Ann Hum Genet. 2009; 73: 314-334.
- Bothwell JE. Pigmented skin lesions in tyrosinase-positive oculocutaneous albinos: a study in black South Africans. Int J dermato. 1997; 36: 831-836.
- Kerr R. Types on albinism in Southern Africa. Proceedings of the 1st international workshop on oculocutaneous albinism in Sub-Saharan Africa. Douala, 2015.
- Aquaron R, Brilliant MH. Tentative genotype-phenotype correlation in Cameroonian and Tanzanian OCA2 patients. Proceedings of the 1st international workshop on oculocutaneous albinism in Sub-Saharan Africa. Douala, 2015.