Review Article | Volume 1 - Issue 2 | Article DOI :
Download PDF
Ma Lin*
The State Key Lab of Pharmaceutical Biotechnology, College of life Sciences, Nanjing University, China
Corresponding Author:
Ma Lin, The State Key Lab of Pharmaceutical Biotechnology, College of life Sciences, Nanjing University,China,
Abstract
RNA silencing is a general surveillance system in plants and animals which could protect hosts from virus infection. However, many virus species survived by expressing a series of proteins named as RNA Silencing Suppressors (RSSRs) to counteract this defense system. This review elaborated the newly discovered RSSRs encoded by virus including the recently discovered polymerase slippage product and some newly-identified RSSRs in mammalian cells. This review will also provide a comprehensive understanding of the role of RSSRs during the virus infection, especially with regard to its newly identified function in epidemic modification in hosts.
Citation
Lin M. Discovery and Roles of Virus-Encoded RNA Silencing Suppressors. SM Virol. 2016; 1(2): 1007.
Introduction
During virus infection in hosts, double strand RNA (dsRNA) as the virus replication intermediates could triggered the RNA silencing mechanisms in hosts [1]. Subsequently small RNA species of 21 to 24 nucleotides are processed by the dicer or dicer-like proteins and then are recruited into the homogenous virus template RNA and subsequently initate the virus template degradation by RISC complex [2]. Although it has been widely reported that RNA interference (RNAi) is a process of eukaryotic posttranscriptional gene silencing that functions in antiviral immunity in plants, nematodes, and insects. However, recent studies provided strong supports that RNAi also plays a role in antiviral mechanism in mammalian cells [3-4]. In order to counteract this defense system, many virus encode diverse RNA silencing supressors (RSSRs) through diverse mechanisms [5]. Here this review summarized several recently discovered RSSRs encoded by diverse virus species in plants and animals. We also elaborated the underlied roles of RSSRs in the host-virus interaction.
Newly Identified RSSRs Encoded by Viruses in Plants and Animals
Recently, many RNA silencing suppressors encoded by different virus species have been reported. By measuring the RNA silencing suppressor activity of proteins encoded by Lettuce Chlorosis Virus (LCV), Kenji Kubota found that P23 possessed the suppressor activity of local RNA silencing. Furthermore, Expression of P23 via agroinfiltration of Nicotiana benthamiana leaves induced local necrosis that increased in severity at elevated temperatures, a novelty given that a direct temperature effect on necrosis severity has not been reported for the other crinivirus VSRs [6-7].
Mingot have shown that a polymerase slippage mechanism in Sweet Potato Feathery Mottle Virus (SPFMV) (genus Potyvirus, family Potyviridae) produces transcripts bringing different ORFs in frame with the upstream polyprotein, thus leading to P1N-PISPO and P3N-PIPO products. The former product PIN-PISPO was found to have the ability to suppress RNA silencing and the WG/ GW motif was required for this interference [7].
Generally the function of binding virus RNA or virus derived small RNAs is the typical character of traditional RSSRs and many of them posess the RNA binding domains. However, it has been found by Kumar that Gemini virus - encoded AC2 have the ability to suppress RNA silencing activity. Meanwhile, unlike many other suppressors, AC2 protein does not bind to siRNA or dsRNA species but interacts with key components of the RNA silencing pathway such as RDR6 and AGO1 [8].
An integral RNA silencing system in plants requires not just the production of RNA silencing in local regions, but also requires the systemic transmission of RNA silencing signal, which is the so-called systemic RNA silencing. In Nguyen’s report, the effects of viral candidate proteins including Rice Grassy Stunt Virus (RGSV) genes s2gp1, s5gp2, and s6gp1 and Rice Ragged Stunt Virus (RRSV) genes s5gp1, s6gp1, s9gp1, and s10gp1were investigated by using the classical Green Fluorescent Protein (GFP) agro infiltration method in Nicotiana benthamiana and it was found that p6 of RRSV displays mild silencing suppressor activity and affects long-range silencing by delaying the systemic silencing signal. In addition, nguyen also identified two new silencing suppressors that displayed mild activity: p2 of RGSV and p9 of RRSV [9].
VSRs have been widely studied for plant viruses and insect viruses, but only a few have been defined for mammalian viruses currently. Recently, Cui L has discovered a novel VSR from Coronaviruses, a group of medically important mammalian viruses including Severe Acute Respiratory Syndrome Corona Virus (SARS-CoV), and showed that the nucleocapsid protein (N protein) of Coronaviruses suppresses RNAi triggered by either short hairpin RNAs or small interfering RNAs in mammalian cells. In addition, the coronavirus N protein could function as a VSR through its double-stranded RNA binding activity [3].
Chen has found that Porcine Reproductive And Respiratory Syndrome Virus (PRRSV) suppressed RNA silencing induced by short-hairpin (sh) RNA, double-strand (ds) RNA and microRNA (miRNA) and down regulated the expression of argonaute protein-2 (Ago-2), which is a key protein of the RNA silencing pathway in animal cells. Furthermore, exogenous introduction of siRNA and shRNA downregulated Dicer or Ago-2 proteins of the cellular RNA silencing apparatus, which, in turn, increased the viral replication and titers [4].
Roles of RSSRs in the Host-Virus Interaction
RSSRs is Required for Virus Infection During its Replication in Hosts
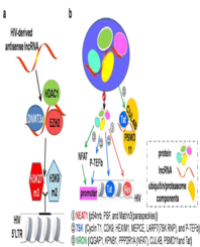
During initial RNA virus infection, the dsRNA replication is required. However, the synthesized viral dsRNA by viral RNA polymerase can also trigger the RNA silencing mechanism and the generation of primary vial siRNA in hosts (Figure 1A).

Figure 1: Roles of RSSRs in the host-virus interaction.
(A) Replication of RNA virus triggers the RNA silencing and counteracts its infection.
(B) RSSRs with the RNA binding domain encoded by virus suppress RNA silencing activity by direct binding with viral dsRNAs or viral small RNAs.
(C) RSSRs lacking the RNA binding domain suppresses RNA silencing activity by interfering with the components of RNA silencing (including dicer, RISC complex, et al.).
(D) RSSRs harrboring the NLS signal alter the expression of virus-resistance-related genes in an epigenetic way.
Then the secondary siRNAs derived from viral RNA templates were produced by host RNA-dependent RNA polymerases (RDRs) [10]. Primary siRNAs are essential but not sufficient for anti-viral defense and the secondary siRNAs were required because loss of RDR1 or RDR6 activity causes increased susceptibility to viral systemic infection in Arabidopsis [11]. It is worth noting that RDR1 from Nicotiana Tabacum (Nt) suppresses the RDR6-mediated antiviral RNAi and enhances viral infection in Nicotiana Benthamiana [12]. Thus, to survive in the hosts in the presence of antiviral RNA silencing mechanism and fulfill the virus replication in hosts, RSSRs are encoded by diverse virus species and exercise their functions including their direct binding to viral dsRNAs or small RNAs and their indirect interference with the normal function of dicer, AGO or RDRPs proteins by binding with them (Figure 1B & C). Actually, virus species lacking of wild type RSSRs failed to successfully infect host cells again.
The Function of RSSRs Could Be Beneficial for the Hosts
Viruses encode RSSRs to facilitate its infection in hosts. However, recent pubulished data indicated that the RSSRs encoded by viruses seem to have effect on triggering antiviral immunity [13-15]. VSRs can be recognized by resistance (R) proteins and triggered its expression by inhibition of Posttranscriptional Gene Silencing (PTGS) in hosts [16]. Also, some other VSRs can elicit Hypersensitive Response (HR) as effectors also triggered [17]. Thus, not only is the involvement of RSSRs essential for virus survival, but also is beneficial for the hosts.
Nucleus Entry of RSSRs Encoded By Virus Inhibits TGS in Hosts
Although the RSSRs are mostly found to have functions of inhibiting RNA silencing in host cytoplasm, it’s also been found that some RSSRs have the NLS signal and could enter the host nucleus, and thus change the TGS by interfering the well-balanced epigenetic events such as RNA-dependent DNA methylation in hosts (Figure 1D). It has been reported that 2b protein encoded by cucumber mosaic virus possesses the NLS signal and locates in the nucleus, and can reduce endogenous DNA Methylation [18]. P19 encoded by Tombusvirus may suppress TGS through binding 21-nt siRNAs or down regulating AGO1 mRNA, which plays a role in methylation of some endogenous loci [19-20].
Conclusion
RNA silencing plays a vital role in the host antiviral defense in plants and animals. To facilitate infection, viruses encode RSSRs to counteract antiviral defense. In conclusion, this review summarized in recent studies newly identified RSSRs encoded by viruses not only in plants but also in animals. During viral infection, antiviral silencing and VSRs have dual effects to maintain the balance between plant growth and virus infection. This review elucidated the underlied mechanisms in the balance between RNA silencing mechanism and its suppression by RSSRs, including the multiple functions of RSSRs encoded by virus species in hosts, revealing the central hub regulators of VSRs in dynamically integrated connections between hosts and viruses.
References
1. Eamens A, Wang MB, Smith NA, Waterhouse PM. RNA silencing in plants: yesterday, today, and tomorrow. Plant Physiol. 2008; 147: 456-468.
2. Andika IB, Kazuyuki Maruyama, Liying Sun, Hideki Kondo, Tetsuo Tamada, Nobuhiro Suzuki. Different Dicer-like protein components required for intracellular and systemic antiviral silencing in Arabidopsis thaliana. Plant Signal Behav. 2015; 10: e1039214.
3. Cui L, Wang H, Ji Y, Yang J, Xu S, Huang X, et al. The Nucleocapsid Protein of Coronaviruses Acts as a Viral Suppressor of RNA Silencing in Mammalian Cells. Journal of virology. 2015; 89: 9029-9043.
4. Chen J, Shi X, Zhang X, Wang L, Luo J, Xing G, et al. Porcine Reproductive and Respiratory Syndrome Virus (PRRSV) Inhibits RNA-Mediated Gene Silencing by Targeting Ago-2. Viruses. 2015; 7: 5539-5552.
5. Roth BM, Pruss GJ, Vance VB. Plant viral suppressors of RNA silencing. Virus research. 2004; 102: 97-108.
6. Kubota K & Ng JC. Lettuce chlorosis virus P23 Suppresses RNA Silencing and Induces Local Necrosis with Increased Severity at Raised Temperatures. Phytopathology. 2016; 106: 653-662.
7. Mingot A, Valli A, Rodamilans B, San León D, Baulcombe DC, García JA, et al. The P1N-PISPO Trans-frame Gene of Sweet Potato Feathery Mottle Potyvirus is produced during Virus Infection and Functions as RNA Silencing Suppressor. Journal of virology. 2016; 90: 3543-3557.
8. Kumar V, Mishra SK, Rahman J, Taneja J, Sundaresan G, Mishra NS, et al. Mungbean yellow mosaic Indian virus encoded AC2 protein suppresses RNA silencing by inhibiting Arabidopsis RDR6 and AGO1 activities. Virology. 2015; 486: 158-172.
9. Nguyen TD, Lacombe S, Bangratz M, Ta HA, Vinh do N, Gantet P, et al. p2 of Rice Grassy Stunt Virus (RGSV) and p6 and p9 of Rice ragged stunt virus (RRSV) isolates from Vietnam exert suppressor activity on the RNA silencing pathway. Virus genes. 2015; 51: 267-275.
10. Fire A, Xu S, Montgomery MK, Kostas SA, Driver SE, Mello CC. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature. 1998; 391: 806-811.
11. Wang X-B, Qingfa Wua, Takao Itoa, Fabrizio Cilloc, Wan-Xiang Lia, Xuemei Chen, et al. RNAi-mediated viral immunity requires amplification of virus-derived siRNAs in Arabidopsis thaliana. Proceedings of the National Academy of Sciences. 2010; 107: 484-489.
12. Ying X-B, Dong L, Zhu H, Duan CG, Du QS, Lv DQ, et al. RNA-dependent RNA polymerase 1 from Nicotiana tabacum suppresses RNA silencing and enhances viral infection in Nicotiana benthamiana. The Plant Cell. 2010; 22: 1358-1372.
13. Soosaar JL, Burch-Smith TM, Dinesh-Kumar SP. Mechanisms of plant resistance to viruses. Nature Reviews Microbiology. 2005; 3: 789-798.
14. Dodds PN, Rathjen JP. Plant immunity: towards an integrated view of plant pathogen interactions. Nature Reviews Genetics. 2010; 11: 539-548.
15. Jones JD, Dangl JL. The plant immune system. Nature. 2006; 444: 323-329.
16. Boccara M, Alexis Sarazin, Odon Thiébeauld, Florence Jay, Olivier Voinnet, Lionel Navarro, et al. The Arabidopsis miR472-RDR6 silencing pathway modulates PAMP-and effector-triggered immunity through the post transcriptional control of disease resistance genes. PLoS Pathog. 2014.
17. Li HW, Lucy AP, Guo HS, Li WX, Ji LH, Wong SM, et al. Strong host resistance targeted against a viral suppressor of the plant gene silencing defence mechanism. The EMBO Journal. 1999; 18: 2683-2691.
18. Duan C-G, Fang YY, Zhou BJ, Zhao JH, Hou WN, Zhu H, et al. Suppression of Arabidopsis ARGONAUTE1-mediated slicing, transgene-induced RNA silencing, and DNA methylation by distinct domains of the Cucumber mosaic virus 2b protein. The Plant Cell. 2012; 24: 259-274.
19. Wang H, Zhang X, Liu J, Kiba T, Woo J, Ojo T, et al. Deep sequencing of small RNAs specifically associated with Arabidopsis AGO1 and AGO4 uncovers new AGO functions. THe plant journal. 2011; 67: 292-304.
20. Wu L, Mao L, Qi Y. Roles of dicer-like and argonaute proteins in TAS-derived small interfering RNA-triggered DNA methylation. Plant physiology. 2012; 160: 990-999.