Research Article | Volume 5 - Issue 1 | Article DOI :
Download PDF
Jasira Jzar1 , Mary Simiyu2 , Joseph Mafurah2*, Joshua Ogendo2 and Anne Osano3
- College of Arts and Sciences, Howard University, Washington DC, USA.
- Department of Crops, Horticulture and Soils, Egerton University, Kenya.
- Department of Natural Sciences, Bowie State University, USA.
Corresponding Author:
Joseph Mafurah, Department of Crops, Horticulture and Soils, Egerton University, Kenya.
Keywords
Chia; Plant growth promotion; Endophytes; Phosphate solubilization; Nitrogen Fixation
Abstract
A total of 9347 fungal and bacterial endophytes were isolated from the roots, stem and leaves of chia plant. Roots harbored more number of fungal endophytes than either stem or leaves whereas stem supported more number of bacterial endophytes than either roots or leaves. The nutritious plant supported more of gram negative compared to gram positive bacterial endophytes. The most common bacteria isolated were Pseudomonas Bacillus, and Cocci. The fungal endophytes isolated from root, stem and leaves of the chia plant showed the presence of Penincillium, Aspergillus, Fusarium, and Macrophomina spps. Dominant fungal endophyte was Aspergillus spp. which was found in all the plant parts instigated. Roots of the plant possessed maximum nitrogen fixers followed by stem and leaves. A proportion of 55% for the bacterial endophytes isolated from the plant chia plant were able to fix nitrogen whereas 25% were able to solubilize phosphorous. The phosphate solubilization efficiency was found to be highest for the Aspergillus spp at 83%.
Introduction
As the human population grows and the climate changes, acquiring and producing all the food needed to support the growing population becomes harder. This problem results in a large-scale food security issue. The increase in farming chia seeds has been one of the ways to combat these food security problems. Chia plant has become a widely grown super-food originally native to Central and Southern America. It is a highly nutrient-dense crop that contains omega-3, omega-6, soluble dietary fiber protein, and phytochemicals [1]. Besides, chia has been shown to have a wide range of health benefits, including prevention against several non communicable diseases, such as obesity, hypertension, Cardiovascular Disease (CVD’s), cancer, and diabetes [1]. The crop is a very versatile and used in many ways in the food industry. As a result of its essential physicochemical and functional properties, it is an excellent thickening agent, gel producer, chelator, foam enhancer, emulsifier, suspension creator, clarifying agent, and rehydrating agent. Consequently, omega-3, protein, fiber, and phenolic compounds can be enriched in new products. Additionally, chia seeds can be added to animal feeds to increase the levels of Alpha-Linolenic Acid (ALA) and lower cholesterol levels [2].
Chia farming faces challenges of pests, diseases and weeds. To combat this problems most farmers use chemical pesticides and herbicides for control. The most common herbicides used are sulfentrazone, oxyfluorfen, s-metolachlor, flumioxazin, diclosulam, and trifluralin [3]. When a large concentration of the pesticides and herbicides are used they can be harmful to the environment and humans or animals and therefore requires alternative means of control [4]. Another common problem Chia farmers have is climate change which occasionally results in severe/ prolonged drought. The best solution to these production challenges is to utilize the plant’s natural systems to resist drought and pest attack through its symbiotic relationship with microbes called endophytes [5].
Endophytes are fungal and bacterial organisms that exist within a plant’s microbiome and have a symbiotic relationships with almost all vascular plants studied. Plant endophytes produce and secrete secondary metabolites and biochemicals that suppress or reduce pathogen growth [6]. In other studies, endophytes induced plant defense mechanisms to protect their hosts [7]. Additionally, to protect their host plants against pathogens, some endophytes have Growth-Promoting Properties (PGPs), which leads to faster growth and stronger plants. These PGP endophytes provide nutrients such as nitrogen, phosphate, and iron, as well as facilitate plant growth and development by growth stimulation [8].
The functions of endophytes when utilized prove to be very useful in agriculture. Some of the most resilient and high-yielding plants contain a wide variety of endophytes, which take on a variety of roles that contribute to the plants’ success [9]. Endophytes can enhance a plant’s ability to grow under unfavorable conditions and could alleviate bacterial and fungal pathogens which will ultimately improve crop yield and production rates [10]. Therefore there meed to replace the harmful pesticides and herbicides that can be detrimental to the environment and human populations by enhancing the plants own specific microbiome. This study aimed at isolation and identification of endophytes in chia plant to promote crop production.
Materials and Methods
The protocol described by [11] was followed
Sample collection
The experiment was carried out at Egerton University Research Field 7 which is located at Njoro Sub-county in Nakuru County, Kenya. It is located at 0°22`S 35°56`E and lies at an altitude of about 2267 m above sea level. The site receives an average annual rainfall of 1800 mm annually with average temperature ranges of between 10.2-22.0°C. The distribution of rainfall is bimodal with long rains between April and August and short rains between October to December. The soils are clay loamy and well drained with a pH of 5.5. Primary and secondary tillage were done obtain fine tilth in April 2023. Well decomposed farm yard manure was applied at the rate of 10 t/ ha and mixed thoroughly with the soil before planting. Certified chia seeds were sown at a spacing of 30 cm by drill. Weeding was done manually by uprooting the weeds as they emerge. No pesticides were applied on the crops throughout the season. The plants were carefully uprooted at flowering stage and bagged in sterile bags and transported to Egerton Biotechnology lab.
Isolation of endophytes
The bacterial endophytes were isolated from the root, stem, and leaves of the collected from a healthy flowering chia Plant. The flowering plant was washed under tap water for 10-15 minutes to remove soil and adhering particles. The plant was air-dried and separated into roots, stems, and leaves. The separated parts were weighed up to one gram then soaked in sterile distilled water and drained. The plant parts were surface-sterilized by dipping them in 70% ethanol for 30 seconds. The leaves and the stem parts were dipped in 4% sodium hypochlorite for 5 minutes and the roots were dipped in 4% sodium hypochlorite for 10 minutes and then treated with 70% ethanol for 30 secs. The separated parts of the leaves, roots, and stem were rinsed five times in sterilized distilled water.
The samples were blot-dried using sterile filter paper and macerated in 10 ml of distilled water by crushing in a mortar using pestle. For each macerated sample, the root, stem, and leaves sample serial dilutions up to 10-5 dilutions were made. One microliter from each dilution of the respective sample was poured into Petri plates labeled from 10-1 to 10-5 containing Nutrient Agar (NA) medium. The plating was done in triplicate for each dilution on Nutrient Agar (NA) for bacterial isolation and on Potato Dextrose (PDA) Agar for fungal isolation. The plates were then incubated at 37°C for 24-48 hours for the isolation of bacterial endophytes and 28°C for two weeks for fungal endophytes. The isolated bacterial and fungal endophytes were maintained as pure cultures on NA and PDA respectively.
Population of bacterial endophytes
Colony-Forming Units (Cfu) of bacterial endophytes were calculated after incubating the bacterial and fungal endophytes at 37°C for two to three days. The Colony-Forming Units (Cfu) of bacterial isolates in roots, stems, and leaves were calculated at the various dilutions using the following equation:
The colonies were counted using the four-quadrat method with the assistance of a digital colony counter.
Morphological and biochemical characterization of bacterial endophytes
Biochemical Characterization of bacterial endophytes from Chia plant. The biochemical traits that had been analyzed during the present investigation includes:
Gram’s reaction was done by adding crystal violet stain is added over the culture. After 1 minute, the stain was poured off, and the excess stain is rinsed with water. Iodine solution was added to cover the smear for 1 minute. The iodine solution was poured off, and the slide is rinsed with running water. A mixture of alcohol and acetone was added then rinsed with water for 5 seconds. The smear was counter-stained with safranin for 1 minute.
Carbohydrate fermentation test: A test tube containing phenol red carbohydrate broth was inoculated with bacteria using inoculation loop. The contents were incubated 37°C for 18 to 24 hours. Yellow colour indicated positive result
Indole test: Five drops of Kovac’s reagent was added to the tube containing the bacteria then results recorded.
Citrate test: Inoculation needle was stabbed into the medium containg bacteria, zigzag the needle back and forth on the surface of the Simmons Citrate Agar slant. The tube containing the slant was incubated for 48 hours. Growth of bacteria is indicative of utilization of citrate.
Catalase test: Using a sterilized inoculating loop, a small amount of bacteria smeared onto the dry slide. A drop of hydrogen peroxide on top of the bacteria. Production of bubbles showed positive result.
Methyl Red - Voges Proskauer test (MR-VP): MR-VP broth, containing peptone, glucose, and a buffer was inoculated with the bacterial sample. After incubation, the methyl red indicator was added. If it turned red, then it was a positive test; if yellow, then it was a negative test.
Hydrogen sulphide production test: Bacteria was inoculated in a tube of sterile nutrient broth. Lead acetate paper strip inserted in the neck of the tube and stoppered well. The inoculated medium was incubated at 37ºC, and examined daily for a blackening of the lower part of the strip.
Gelatin test: a needle containing gelatin was stabbed deep all the way to the bottom of the tube containing bacteria. It was incubated at 25ºC for one week. Gelatin hydrolysis was indicated by clear zones around gelatinase (liquefaction).
Starch: Five to ten drops of iodine solution was added directly to the bacterial solution. A blue-black color indicated presence of starch.
Capsular staining: thin smears of bacterial culture were prepared on a microscope slide. The smear was air dried. Crystal violet (1%) was applied and allowed it to remain on the slide for 2 minute then gently washed off with 20% copper sulfate.
Motility test: Sterile needle was used to pick a well-isolated colony and stabbed the nutrient broth to within 1 cm of the bottom of the tube. It was incubated at 35°C for 18 hours until until growth was evident.
Morphological characterization of fungal endophytes
The isolated fungal endophytes were identified based on different morphological features such as colony characterization, growth of fungi, the color of the colony (front and reverse), size, and the shape of conidiophores and conidia under microscopic observation using lactophenol blue staining.
Nitrogen fixation by bacterial endophytes
The nitrogen-fixing ability of the bacterial endophytes was detected by inoculating the isolated pure endophytic bacterial test cultures on the Petri plates containing Jensen’s media at 37°C for 5 days.
Phosphate solubilization of bacterial and fungal endophytes
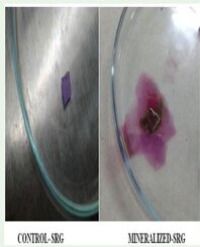
The phosphate solubilizing ability of the bacterial and fungal endophytes was detected by spot inoculating pure isolated endophytic bacterial and fungal cultures separately on Pikovskaya’s medium and incubated at 37°C for three days and 28°C for seven days respectively along with the control plates. The uninoculated plates served as the control. All the inoculations were done in triplicate. Phosphate solubilization by the endophytic bacterial and fungal cultures was tested by their ability to solubilize inorganic phosphate. The Phosphate Solubilization Efficiency (PSE) (Nguyen et al. 1992) will be determined by:
Results
Isolation of bacterial and fungal endophytes
A total of 9,347 bacterial and fungal colonies were isolated from the roots, stem and leaves Chia plant.The bacterial endophytes were higher at 9330 colonies while fungal colonies were 17.
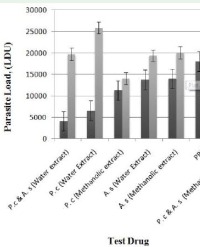
The highest isolation of bacterial endophytes was realized at a dilution of 10-5 with 3264 colonies while the lowest was at 10-3 with 922 colonies.The stem had the highest bacterial colonies at 4737 while the root had the lowest at 1623 (Table 1).The most dominant bacteria were Pseudomnas, Bacillus and Cocci.
Fungal endophytes were isolated in high amount at a dilution of 10-4 with 5 fungal colonies while 10-4 was the lowest with one colony. Additionally, more fungi was isolated in the root with 9 colonies while lowest isolation was from the leaf with two colonies (Table 1). Out of 9347 bacterial and fungal endophytes only 28 bacterial endophytes and 6 fungal endophytes had the ability to grow under laboratory conditions.
Table 1: Number of bacterial and fungal colonies in each dilution and plant parts.
| Dilution/ |
10-Jan |
10-Feb |
10-Mar |
10-Apr |
10-May |
| Part |
| |
Bct |
Fu |
Bct |
Fu |
Bct |
Fu |
Bct |
Fu |
Bct |
Fu |
| Leaf |
1076 |
1 |
808 |
0 |
1 |
0 |
1 |
1 |
1084 |
0 |
| Stem |
1 |
1 |
1272 |
0 |
916 |
1 |
1528 |
2 |
1020 |
2 |
| Root |
1 |
2 |
456 |
0 |
5 |
2 |
1 |
2 |
1160 |
2 |
| Total |
1078 |
4 |
2536 |
1 |
8922 |
3 |
1530 |
5 |
3264 |
4 |
Key: Bct: Bacteria; Fu: Fungi
Morphological characterization of bacterial endophytes
The results showed that out of 9330 bacterial endophytes isolated from roots, stem and leaves of the Chia plant, 90.12% of the bacteria had round colony shape, 85.03% had entire margins and 70% had convex elevation. The color of the colony which was most pronounced among the bacteria isolated from the leaf at dilution across all dilutions was white. The roots and stem had white, cream and yellow colonies.
Biochemical characterization of bacterial endophytes from Chia plant
Out of the 10 bacteria isolated from roots that could grow on NA, 85% were Gram negative (Plate 1).

Plate 1 Gram positive bacteria ( x400).
All were positive for citrate and catalase, while 7 were positive for VP, gelatin, motility and starch and only 2 of them could ferment three sugars and most of them were MR and indole negative.
From the stem, out of the 10 isolated bacteria, all were gram negative (Plate 2).

Plate 2 Gram negative bacteria ( x400).
Half were rod shaped and the other half was capsule shaped. only 3 were citrate and catalase positive and most of them were negative for VP, gelati and starch while all were found to be negative for MR, indole and motility.
The leaves had 8 bacteria growing on plates and all were gram negative (Plate 2) and rod shaped. Six were citrate, VP, and starch positive and only two of them could ferment three sugars i.e. lactose, sucrose and glucose while only one of them was catalase positive. All were negative for gelatin, MR and indole and motility.
Morphological Characterization of fungal endophytes
The endophytic fungi isolated from healthy roots of the plant Chia included Aspergillus spp, Macrophomina spp and Fusarium spp. The stem had Fusarium spp, while the leaves were dominated with Penicillium spp (Figure 1).

Figure 1 Plates depicting morphological characteristics of the isolated fungi:a) and b) front view and pycnidiohores of Penicillium spp, c) and d) front view and pycnidiohores of Fusarium spp,e) and f) front view and pycnidiophores of Aspergillus spp,g) and h) front view and pycnidiohores of Macrophomina spp (Pycnidiophores were observed at x400).
Plant growth promoting traits of the isolated endophytes
55.83% of the bacterial endophytes isolated from the Chia plant were able to fix nitrogen.Roots of the plant possessed maximum nitrogen fixers followed by stem and leaves, whereas only one fungi was able to soluibilize phosphorous.
Nitrogen fixation by bacterial endophytes: Out of 10 bacterial endophytes isolated from roots of the plant Chia plant 90% were able to fix nitrogen whereas out of 10 bacterial endophytes isolated from the stem 40% were able to fix nitrogen and out of 8 bacterial endophytes isolated from leaves 37.5% were able to fix nitrogen. In total 55.83% of the bacterial endophytes isolated from the Chia plant were able to fix nitrogen. Roots of the plant possessed maximum nitrogen fixers followed by stem and leaves.
Phosphate solubilization by fungal endophytes: Only one fungi isolated from the roots of Chia plant (Aspergillus spp) was able to solubilize phosphates. The solubilization efficiency for the endophyte was found to be 83%.
Discussion
Research and production of chia seed has largely been neglected in Kenya and its not widely grown in most parts of the country despite its huge nutritional importance. The crop has the potential to become an important substitute for most crops that are not drought tolerant to improve food security, ensure production resilience and maintain the health of consumers. This study represents the first of its kind on chia plant and is an important first step for further such studies towards crop improvement. Results detected several generra of bacteria and fungi genera from one location. For fungi, genera such as Aspergillus, Fusarium, Macrophomina and Penicillium were among the dominant groups in this study. This is in agreement with [12] who found out that almost all vascular plant species investigated to date have been found to be home for endophytic microorganisms. This is because the plants provide a conducive environment for the growth and reproduction of endophytic bacteria and fungi. Endophytes and mycorrhizal fungi are considered beneficial symbionts which promote plant growth, confer enhanced resistance to various pathogens and pests, and improve survival under unfavorable environmental conditions [13].
This study revealed that bacterial colonies were recovered in serial dilutions of different plant parts, including the stem, leaf, and roots. Higher abundance of bacterial colonies were recovered at 10-5 with the highest density detected in the stem, followed by the leaves and roots, suggesting a differential distribution pattern within the plant. This result is in agreement with the research done [11]. Most of the bacteria isolated from this study were gram negative and positive for biochemical reactions. A proportion of 55% demonstrated the ability to fix nitrogen. Earlier studies have revealed a predominance of gram negative bacteria in the tissues of various plants [14]. The research conducted by [15], managed to isolate 667 bacteria in chia out of which thirteen strains showed all the tested PGP activities. Furthermore, two strains were found to show highest salt tolerance. Similar results were reported in Amaranthus spinosus where 62.75% of the bacterial endophytes isolated from the plant were able to fix nitrogen [11].
The fungal genera that were identified were Aspergillus spp, Fusarium spp, Macrophomina spp, and Penicillium spp. The genus Aspergillus was characterized by the formation of flask-shaped or cylindrical phialides either in a single or double series on the surface of a vesicle at the apex of a conidiophore Conidia are deciduous and globose, oblong to elliptical, and present various colors but commonly green and yellow [16]. The three species of Aspergillus spp identified showed yellow-greenish features coupled with the round-like vesicles and the spherical light green conidia and Green-yellowish round nodes of the conidiophores.
Aspergillus showed to have phosphorus solubilizing capabilities with a Phosphate Solubilization Efficiency (PSE) of 8.3% [17]. This capability shows the relationship between the endophyte and the chia plant is for the endophyte to solubilize phosphate for the plant to utilize, which promotes plant growth.
Some of the putative fungal endophytes isolated in this study belong to species that are known pathogens and would not be considered for future studies without eliminating that possibility through further molecular analysis. Future studies will focus on testing these endophytes for their ability to increase abiotic/biotic stress tolerance in chia plant. Additional isolation of endophytes from chia plant at different times during their growth cycle and from relatively dry area could also be beneficial for expanding this endophyte collection. This endophyte collection will allow us to begin evaluating their potential for increasing agricultural production in a sustainable, and environmentally friendly manner.
Conclusion
This study demonstrated that chia plant supports wide range of bacteria and fungi that are symbiotic in the roots, stem and leaves. The nutrient-dense plant harbored more bacteria in the microbiome as compared to the number of fungi. Most of the isolated bacteria were positive to the biochemical activities tested. Besides, a large percentage of bacteria had the ability to fix nitrogen as per the Jensens experiment. Some of the fungi isolated have been found to be pathogenic while others are endophytic. One fungi was able to solubilize phosphates and this implies that this endophyte can be incorporated in the soil with the seed to improve phosphorous uptake in the soil. The presence of nitrogen fixing and phosphorous solubilizing endophytes gives a positive insight for future research to produce chia seed at low cost and limit the use of synthetic fertilizers that are pollute the soil and and hazardous to man. Further research is however recommended to determine the functions of the other isolated endophytes.
Acknowledgments
We would like to express our sincere gratitude to the U.S. National Science Foundation Research Experiences for Undergraduates (REU) for funding the project. The Global Research Experience for Undergraduate students (REU) program is collaboration between Bowie State and Egerton University. It at training Globally Engaged Undergraduates in Food Security Research in Kenya: A Focus on Tropical Food Crops, as the Key to Feeding the World (Award No. 2150005).
References
1. Suri S, Passi JS, Goyat, J. Chia seed (Salvia hispanica L.): A new age functional food. 4th International Conference on Recent Innovations in Science Engineering and Management. 2016; 4(3): 286-299.
2. Ullah R, Nadeem M, Khalique A, Imran M, Mehmood S, Javid A, et al. Nutritional and therapeutic perspectives of Chia (Salvia hispanica L.): A review. J Food Sci Technol. 2016; 53: 1750-1758.
3. Maciel CDG, Vidal LHI, Jadoski SO, Iuchemin CEL, Helvig E, da Silva AAP, et al. Selectivity of herbicides applied in pre-emergence in chia crop. Revista Brasileira de Ciências Agrárias - Brazilian Journal of Agricultural Sciences. 2019; 14: 1-7.
4. Duke SO. Overview of herbicide mechanisms of action. Environmental Health Perspectives.1990; 87: 263-271.
5. de Oliveira Costa LE, de Queiroz MV, Borges AC, de Moraes CA, de Araújo EF. Isolation and characterization of endophytic bacteria isolated from the leaves of the common bean (Phaseolus vulgaris). Braz J Microbiol. 2012; 43: 1562-1575.
6. Kottb M, Gigolashvili T, Großkinsky DK, Piechulla B. Trichoderma volatiles effecting arabidopsis: From inhibition to protection against phytopathogenic fungi. Front Microbiol. 2015; 6: 995.
7. Shoresh M, Harman GE, Mastouri F. Induced systemic resistance and plant responses to fungal biocontrol agents. Annu Rev Phytopathol. 2010; 48: 21-43.
8. Saleem M, Arshad M, Hussain S, Bhatti AS. Perspective of Plant Growth Promoting Rhizobacteria (PGPR) containing ACC deaminase in stress agriculture. J Ind Microbiol Biotechnol. 2007; 34: 635-48.
9. Nair DN, Padmavathy S. Impact of endophytic microorganisms on plants, environment and humans. Scientific World Journal. 2014; 2014: 250693.
10. Narayanan MM, Ahmad N, Shivanand P, Metali F. The role of endophytes in combating fungal- and bacterial-induced stress in plants. Molecules. 2022; 27: 6549.
11. Sharma S, Roy S. Isolation and Identification of a novel endophyte from a plant Amaranthus spinosus. International Journal of Current Microbiology and Applied Sciences. 2015; 4: 785-798.
12. Firakova S, Sturdikova M, Muckova M. Bioactive secondary metabolites produced by microorganisms associated with plants. Biologia. 2017; 62: 251–257.
13. Rodriguez RJ, Redman RS, Henson JM. The role of fungal symbioses in the adaptation of plants to high stress environments. Mitigation and Adaptation Strategies for Global Change. 2004; 9: 261-272.
14. Elbeltagy A, Nishioka K, Suzuki H, Sato T, SatoYI, Morisaki H, et al. Isolation and characterization of endophytic bacteria from wild and traditionally cultivated rice varieties. Soil Science and Plant Nutrition. 2000; 46: 617-629.
15. Yañez-Yazlle MF, Romano-Armada N, Rajal VB and Irazusta VP. Amelioration of saline stress on chia (Salvia hispanica L.) seedlings inoculated with halotolerant plant growth-promoting bacteria isolated from hypersaline environments. Front. Agron.2021; 3: 665798.
16. Gouda S, Das G, Sen SK, Shin HS, Patra JK. Endophytes: A treasure house of bioactive compounds of medicinal importance. Front Microbiol. 2016; 7: 1538.
17. Raper KB, Fennell DI. The genus aspergillus. Philadelphia. Williams and Wilkins. 1965: 686.
Citation
Jzar J, Simiyu M, Mafurah J, Ogendo J, Osano A. (2024) Characterization of Endophyte Microbiome Diversity in Chia Plant (Salvia hispanica L.). Ann Appl Microbiol Biotechnol J 5: 5.