Review Article | Volume 1 - Issue 1 | Article DOI :
Download PDF
Xing Ye*and Yuan-yuan Tian
Key Laboratory of Tropical and Subtropical Fisheries Resource Application and Cultivation, China Ministry of Agriculture; Pearl River Fisheries Research Institute, Chinese Academy of Fishery Sciences, Guangzhou, China
Corresponding Author:
Ye Xing, China Ministry of Agriculture, Chinese Academy of Fishery Sciences, China
Keywords
Grass Carp Reovirus; Molecular Type; Immunogen; Vaccine
Abstract
Grass carp reovirus, GCRV, belongs to the genus Aquareovirus (AQRV). It is the most virulent species of AQRV, and infection by GCRV causes hemorrhagic disease in grass carp. A new strain, GCRV-GD108, was found in China. Significant differences were found between GCRV-GD108 and GCRV as well as between GCRVGD108 and other known AQRVs. Moreover, similarities were found between GCRV-GD108 and Orthoreovirus (ORV), suggesting a closer evolutionary relationship between GCRV-GD108 and ORV than between GCRVGD108 and the known AQRVs. The discovery of different virus molecular types of GCRV indicates the importance of molecular diagnosis and the development of a specific vaccine. Vaccines have been developed that include inactivated tissue vaccines, inactivated cell vaccines, and attenuated viral vaccines. Great efforts have been made in recent years to investigate immunogen for the preparation of genetically engineered vaccines, which are expected to provide protection for the cultured grass carp.
Keywords
Grass Carp Reovirus; Molecular Type; Immunogen; Vaccine
CITATION
Ye X and Tian Y. Advances in GCRV Research: Virus Molecular Type and Immunogen. SM Virol. 2016; 1(1): 1003.
Introduction
The grass carp (Ctenopharyngodon idellus) is one of the most important aquaculture species in China. However, grass carp are vulnerable to pathogens such as GCRV, which can cause severe hemorrhage, resulting in a high mortality rate. GCRV was first isolated in China in 1983[1] and has been recognized by the International Committee on Taxonomy of Viruses (ICTVs) as a species belonging to the genus Aquareovirus (AQRV) [2]. GCRV is icosahedral in symmetry with a diameter ranging between 70 and 90 nm. Its genome consists of 11 segments of dsRNA that are packaged into a two-shell capsid without an envelope. Through years of effort, great progress has been achieved in the study of the biological characteristics, invasion, replication, and assembly mechanism of GCRV [3-5], as well as in vaccine development [6-8]. In recent years, different molecular types of GCRV have been discovered, indicating the importance of the preparation of specific vaccines in order to effectively prevent hemorrhagic disease.
Discovery of Various Grass Carp Reoviruses
The reovirus that infects fish belongs to AQRV, which usually possesses a genome of 11 dsRNA segments. Seven AQRV genetic groups were established (designated A to G) in genus AQRV, and GCRV belongs to AQRV-C [9,10]. GCRV-873 is the representative strain of GCRV, and it is the f irst strain for which the entire genomic sequence was obtained [11]. In 2009, our group isolated a strain, GCRV-GD108, from diseased grass carp in Guangdong province, China, and proved that it is the pathogen responsible for the hemorrhage disease [12]. We obtained the entire sequence of the genome segment. Significant differences were found between GCRV-GD108 and GCRV-873, as well as between GCRV-GD108 and other known AQRVs [12]. Further investigations and comparisons of virus strains in Guangdong, Fujian, Hunan, and other provinces in China demonstrated that they share high molecular homology with GCRV-GD108, indicating that GCRV-GD108 is a representative strain in southern China [13].
GCRV-GD108 could be classified as belonging to the genus AQRV, according to criteria of demarcation [12]. However, it also exhibited a number of differences in relation to known species of AQRV. These include the following: 1) The VP5 of GCRV-GD108 did not exhibit high homology to AQRVs (24–25%), whereas among different AQRVs, VP5 exhibited high sequence identity (57 92%), suggesting that GCRV-GD108 may be a new species of AQRV;2) the GCRV-GD108 sigma 1-like protein (encoded by S7) was absent in AQRVs; 3)the S10 gene of GCRV-GD108 encoded a protein that was not homologous to the VP7 protein of AQRVs, and no homology to VP7 was found in GCRV-GD108[12].Furthermore, GCRV-GD108 displayed a closer evolutionary relationship to Orthoreovirus (ORV) than to the known species of AQRV. For example, like the ORVs, GCRV GD108 caused no syncytia, where as syncytia is a typical cytopathic effect of infection in a majority of AQRVs; the sigma-1-like protein encoded by S7 of GCRV-GD108, which was absent in AQRVs, was homologous to mammalian orthoreoviruses (MRVs).
To date, the full genomic sequences of at least 11 strains of GCRV have been obtained [14-18]. Most of these were isolated in China, except for the golden shiner reovirus (GSRV) and the American grass carp reovirus (AGCRV), which were isolated in the USA [19,20]. T hese strains could be divided into three groups [17]: The first group includes GCRV-GD108 and most strains of GCRV, the second group includes GCRV-873, and the third group contains only one member, GCRV-104(HGDRV) [18]. A phylogenetic tree has been constructed based on the sequences of the RdRp of AQRVs and ORVs in Figure 1.

Figure 1: Phylogenetic tree built based on the putative amino acid sequence of RNA-dependent RNA polymerase, VP2 of partial members of Aquareovirus and Orthoreovirus, using Maximum parsimony method (MEGA5.03). Scale bar indicates the number of substitutions per aligned a position giving rise to the phylogram. Orthoreovirus: ARV (Avian orthoreoviruses, GenBank accession no. ACH72476); NBV (Nelson Bay orthoreoviruses, AEQ49381); BroV tentative Orthoreovirus species, ACU68602; BRV (Baboon orthoreovirus, strain Baboon reovirus, AEK86190; MRV (Mammalian orthoreovirus, strain Type 1 Lang, AAA47234); PRV (tentative Orthoreovirus species, strain Reovirus Salmo/GP-2010/NOR, GU994015). Aquareovirus: AQRV-A (strain Scophthalmus maximus reovirus, ADZ31977); AQRV-C (strain grass carp reovrius 873, AAG10436.1); AQRV-G (strain AGCRV PB01-155, ABV01040); tentative Aquareovirus species, strain GCRV GD108, ADT79734.1; tentative Aquareovirus species, strain GCRV-104, AFG73673.
A more detailed phylogenetic tree based on AQRVs and ORVs structural proteins can be referred to in the work of Nibert and Duncan [14]. The virus genotypes in diseased grass carp isolated in southern China by our group were similar to that of GCRV-GD108, and a low detection rate of GCRV-873 and GCRV-104 (HGDRV) was found by several other groups [private communication]. Whether these two genotypes, GCRV-873 and GCRV-104 (HGDRV), are now pathogenic to grass carp remains unknown. Regardless, the existence of various genotypes of GCRV suggests the importance of molecular diagnosis and the development of a specific vaccine.
Development of GCRV Vaccines
Because of the significant importance of grass carp aquaculture industry in China, great efforts have been made in the development of vaccines to prevent grass carp hemorrhage disease. In the early 1960s, an inactivated tissue vaccine was developed using the formalin-inactivated method to treat the diseased fish tissues, and it demonstrated efficacy in the prevention of hemorrhagic disease [8]. However, it has limitations, which include finite sources and regional variability. In the 1980s, a grass carp kidney cell line was successfully established, and inactivated viral vaccines were subsequently prepared [6-8]. In the late 1990s, an attenuated viral vaccine for grass carp was developed, which showed a 100% survival rate and immune protection rate [7]. This attenuated viral vaccine had obtained its approval number as a veterinary drug product in 2011 and is the first one permitted for use in Chinese aquaculture.
Compared with inactivated and attenuated viral vaccines, a genetically engineered vaccine has the advantages of low cost, no pathogencity, etc. However, selection of the appropriate antigen is the prerequisite for the preparation of a genetically engineered vaccine. T herefore, great efforts have been made in recent years to investigate the immunogen of GCRV for vaccine design.
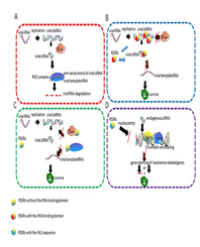
The GCRV capsid structure has two layers: a turreted core enclosed by an outer capsid shell [21]. The core consists of three proteins (VP1, VP3, and VP6), while the outer shell is constituted by VP5/VP7 dimers. The immunogenicity of the VP5 and VP7 proteins of GCRV has been well demonstrated [22-24]. Recently, Hao et al. further demonstrated that VP7 proteins induce a stronger immune response in grass carp than the other GCRV structural proteins, implying that the VP7 protein could be used as a preferred immunogen for vaccine design [25].
In contrast to other AQVRs, 11 dsRNA segments of GCRV GD108 encoded 11 proteins instead of 12 proteins. No homology to the VP7 protein of AQRVs was found in GCRV-GD108, while a homologous protein of MRV sigma-1-like protein encoded by the S7 gene of GCRV-GD108 was found to exist [12]. Our group demonstrated that this sigma-1-like protein possessed strong immunogenicity and protection efficiency, and it is supposed to be a cell attachment protein. Further investigation of its function is ongoing [unpublished data]. The VP4 protein encoded by the M6 gene of GCRV-GD108 is supposed to be a major outer capsid protein, and we have also demonstrated that a polyclonal antibody against recombinant VP4 protein could prevent viral infection efficiently and that recombinant VP4 protein provides good protection for grass carp, with a protection rate as high as 82% [26]. A genetically engineered vaccine of GCRV-GD108 that can provide effective protection has recently been developed by our group, and we are now applying for field trial. This vaccine is expected to be applied in grass carp aquaculture in the near future.
References
1. Chen BS, Jiang Y. Morphological and physico-chemical characterization of the hemorrhagic virus of grass carp. KexueTongbao. 1984; 29: 832-835.
2. Francki RIB, Fauquet CM, Knudson DL, Brown F. Classification and nomenclature of viruses. Fifth report of the International Committee on Taxonomy of Viruses. Arch. Viorl. 1991; 2: 186-192.
3. Fang Q, Ke LH, Cai YQ. Growth characterization and high titre culture of GCHV. Virol. Sin. 1989; 3: 314-339.
4. Zhang X, Jin L, Fang Q, Hui WH, Zhou ZH. 3.3 A cryo-EM structure of a nonenveloped virus reveals a priming mechanism for cell entry. Cell. 2010; 141: 472-482.
5. Cheng L, Fang Q, Shah S, Atanasov IC, Zhou ZH. Subnanometer-resolution structures of the grass carp reovirus core and virion. J Mol Biol. 2008; 382: 213-222.
6. Yang XL, Xia C, Zuo WG. Inactive vaccine for hemorrhage of grass carp: comparison of immunogenicity and immunizing dose between two strains. Journal of Fisheries of China.1989; 13: 138-144.
7. Xu SY, Li HL, Deng GC, Jiang L. The preparation and immune effect of attenuated live vaccine obtained through cell culture for hemorrhage of grass carp. Journal of Fisheries of China.1994; 18:110-117.
8. Zeng L, Yang X, He L, Ai X, He J, Zuo W. Batch processing technology of cell cultured killed vaccine against the hemorrhage of grass carp. J Fishery Sci China. 1998; 5: 62-67.
9. Rangel AA, Rockemann DD, Hetrick FM, Samal SK. Identification of grass carp haemorrhage virus as a new genogroup of aquareovirus. J Gen Virol. 1999; 80: 2399-2402.
10. Attoui H, Mertens PPC, Becnel J, Belaganahalli S, Bergoin M, et al. Reoviridae. In: King AMQ, Adams MJ, Carstens EB, Lefkowitz EJ, editors. Virus Taxonomy: Ninth Report of the International Committee on Taxonomy of Viruses. San Diego: Elsevier/Academic Press. 2012; 541-637.
11. Qiu T, Lu RH, Zhang J, Zhu ZY. Molecular characterization and expression of the M6 gene of grass carp hemorrhage virus (GCHV), an aquareovirus. Arch Virol. 2001; 146: 1391-1397.
12. Ye X, Tian YY, Deng GC, Chi YY, Jiang XY. Complete genomic sequence of a reovirus isolated from grass carp in China. Virus Res. 2012; 163: 275-283.
13. Chi YY, Tian YY, Ye X, Deng GC, Li J, Wang HJ. Molecular properties of grass carp reovirus in southern China and establishment of a duplex PCR detection method. Chin J Virol. 2011; 27: 358-364.
14. Nibert ML, Duncan R. Bioinformatics of recent aqua- and orthoreovirus isolates from fish: evolutionary gain or loss of FAST and fiber proteins and taxonomic implications. PLoS One. 2013; 8: e68607.
15. Zhang Q Y, Gui J F. Virus genomes and virus-host interactions in aquaculture animals. Sci China Life Sci. 2015; 58: 156-169.
16. Wang Q, Zeng W, Liu C, Zhang C, Wang Y, Shi C, et al. Complete genome sequence of a reovirus isolated from grass carp, indicating different genotypes of GCRV in China. J Virol. 2012; 86: 12466.
17. Pei C, Ke F, Chen ZY, Zhang QY. Complete genome sequence and comparative analysis of grass carp reovirus strain 109 (GCReV-109) with other grass carp reovirus strains reveals no significant correlation with regional distribution. Arch Virol. 2014; 159: 2435-2440.
18. Fan Y, Rao S, Zeng L, Ma J, Zhou Y, Xu J, et al. Identification and genomic characterization of a novel fish reovirus, Hubei grass carp disease reovirus, isolated in 2009 in China. J Gen Virol. 2013; 94: 2266-2277.
19. Plumb JA, Bowser PR, Grizzle JM, Mitchell AJ. Fish viruses: a double stranded RNA icosahedral virus from a North American cyprinid. Journal of the Fisheries Research Board of Canada. 1979; 36: 1390-1394.
20. Mohd Jaafar F, Goodwin AE, Belhouchet M, Merry G, Fang Q, Cantaloube JF, et al. Complete characterisation of the American grass carp reovirus genome (genus Aquareovirus: family Reoviridae) reveals an evolutionary link between aquareoviruses and coltiviruses. Virology. 2008; 373: 310-321.
21. Zhu J, Cheng L, Fang Q, Zhou ZH, Honig B. Building and refining protein models within cryo-electron microscopy density maps based on homology modeling and multiscale structure refinement. J Mol Biol. 2010; 397: 835-51.
22. Zhang LL, Lei CF, Fan C, Fang Q, Jin-yu SHEN, Gui-jie HAO. Expression of outer capsid protein VP5 of Grass carp reovirus and analysis of its immunogenicity. Virol Sin. 2009; 24: 545-551.
23. He Y, Xu H, Yang Q, Xu D, Lu L. The use of an in vitro micro neutralization assay to evaluate the potential of recombinant VP5 protein as an antigen for vaccinating against grass carp reovirus. Virol J. 2011; 8: 132.
24. Shao L, Sun X, Fang Q. Antibodies against outer-capsid proteins of grass carp reovirus expressed in E. coli is capable of neutralizing viral infectivity. Virol J. 2011; 8: 347.
25. Hao GJ, Yuan XM, Pan XY, Xu Y, Yao JB, Lin LY, et al. The construction of pEGFP-n1-vp7 containing GCRV VP7 gene and studies on its immune efficacy. Acta Hydrobiologica Sinica. 2015; 39: 751-757.
26. Tian Y, Ye X, Zhang L, Deng G, Bai Y. Development of a novel candidate subunit vaccine against grass carp reovirus Guangdong strain (GCRV GD108). Fish Shellfish Immunology. 2013; 35: 351-356.